
Anatomy of a Data Product — Part Three
This is an article in a series about building data products with Tag.bio.
To begin the series, check out Part One, which outlines the reason for and definition of a data product, along with key concepts and terms. To access the data & codebase to follow along with these examples, see Part Two.
Here, in Part Three, we will introduce the low-code config layer, which ingests and transforms source data into a versioned, well-modeled data snapshot.
Now that we are introducing code, it’s a good time to link to our syntax reference guide — here’s the entry page for config syntax. Almost all code shown will use JSON. It’s possible to alternatively use YAML, if you prefer it.
What the config does
Key words in bold will be explained in more detail below.
The config:
- Loads data from one or more tabular sources — e.g. CSV, TSV, SQL, etc.
- Defines rules for parsing the rows of each table, and for joining multiple tables.
- Specifies which type of entities will be the primary focus of the data model.
- Incorporates parsers — functions which will extract, transform, and load data from the tables as collections of variables into the data model.
Note — only a fraction of config functionality will be shown in this simple example.
How to find the config
The conventional location for the config within a data product codebase is config/. Typically there is one file within that folder — called config.json — which controls the process.

Top-level config attributes
For this example, we will only discuss the config attributes required for fc-iris-demo. For a more comprehensive list, see the syntax reference guide.
- data_dictionary — specifies a file which will contain the data dictionary after the data is loaded and modeled.
- entity_table — a table object which specifies the source data via the table attribute, unique_keys to define entities, and parsers which will do the work of loading and structuring the data.
That’s it — for this simple case. Additional config attributes not shown for this example are other_tables, foreign_keys, null_indicators, joins, and SQL connectivity & query options.
The importance of entities
A data product is built around a single type of entity — e.g. patient, sample, customer, event — in order for questions to be asked about sets (cohorts) of those entities.
Entities are defined by a distinct combination of values over all columns specified by unique_keys in the entity_table. In the data model produced by the config, the entities will represent the “rows”.
For the fc-iris-demo example, each entity represents an individual flower represented by distinct value in the observation column of the source data.

Now let’s drill into parsers
Information about entities is collected and structured in the data model as collections of variables. While entities represent “rows” within the data model, variables represent “columns” — and variables are grouped into higher-order collections.
Variables are created by parsers. Here’s an example:

This parser will load the values from the sepal_length column in the source data table and associate them with entities in the “Sepal length” variable. That variable will be added to the “Sepal” collection for future use.

Note the first attribute — parser_type. The simplest parser_types are “numeric” and “categorical”, although there are 50+ parser_types which can perform a useful variety of complex operations for loading data from rows and columns.
Here’s an example of a “categorical” parser_type — it will load values from the species column in the source data table and associate them with entities in three dynamically-named variables in the “Species” collection.

Each generated categorical variable in the “Species” collection will correspond to one of the three distinct values of species, as a one-hot (AKA dummy) encoding of that data.
- Iris-setosa
- Iris-versicolor
- Iris-virginica
To put it another way, each of these categorical variables in the “Species” collection represents the set of entities belonging to each species.

Data harmonization
Note in both parser examples above, and in all parsers for fc-iris-demo, the resulting variable and collection names have spaces instead of underscores, and begin with capital letters.
This is an important transformation of the source data. Future applications built with this data product — either protocols, or extracted data frames in R & Python — will be able to use these interpretable, harmonized names as fields & features.
Harmonization significantly increases the robustness of data products — e.g. when converting a data product to use a different data source, or when external code accesses multiple data products built from disparate underlying sources — applications built with harmonized data products will still work!
Managing complexity
The fc-iris-demo example is simplistic and does not perfectly demonstrate the complexity of modeling data from multiple sources of data with many columns.
In practice, Tag.bio developers will not include all table objects and parser arrays inline within a single config file — otherwise that file could get really long. Instead, we slice up the config content into multiple, modular files and reference those files from within the config.
In other words, all JSON objects and arrays in the config can alternatively be references to other files which contain those objects and arrays.
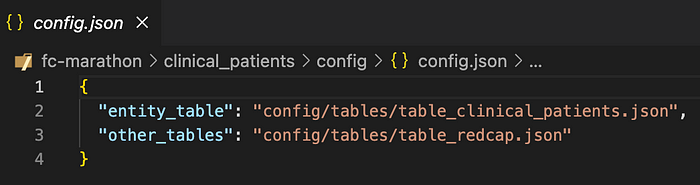
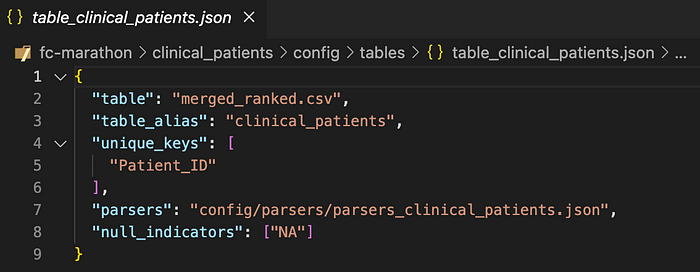
In addition, the Tag.bio framework provides a utility to infer and auto-generate basic parsers from any source data table, automatically seeding the config with collateral as you develop.
Putting it all together
The Tag.bio framework will utilize the config instructions to load and model the source dataset(s) into a data snapshot, which is stored to a file (typically) outside of the code repository. In the case of this example, however, the snapshot is written to the file _data/archive.ser, shown in the screenshot below.
If the config specifies a data_dictionary, then a TSV file will be auto generated which describes the data model in the snapshot.

What’s next?
Part Four in this series will show how low-code protocols define API methods for invoking useful, domain-driven, queries, algorithms and visualizations.
